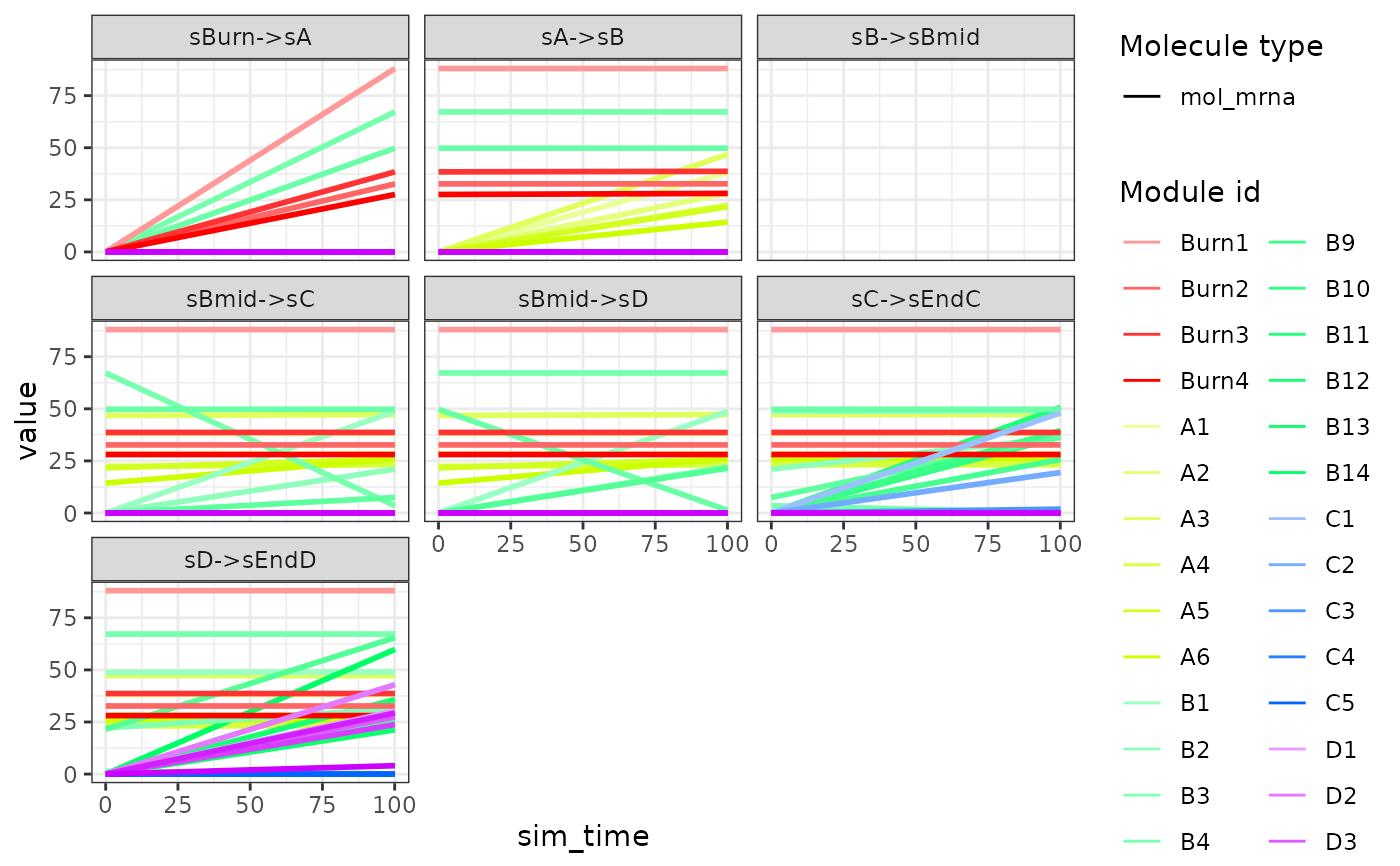
Visualise the expression of the gold standard over simulation time
Source:R/plotting.R
plot_gold_expression.RdVisualise the expression of the gold standard over simulation time
plot_gold_expression(
model,
what = c("mol_premrna", "mol_mrna", "mol_protein"),
label_changing = TRUE
)Arguments
- model
A dyngen intermediary model for which the simulations have been run with
generate_gold_standard().- what
Which molecule types to visualise.
- label_changing
Whether or not to add a label next to changing molecules.
Value
A ggplot2 object.
Examples
data("example_model")
plot_gold_expression(example_model, what = "mol_mrna", label_changing = FALSE)
#> `geom_line()`: Each group consists of only one observation.
#> ℹ Do you need to adjust the group aesthetic?